Simulation of Pumas Models
Simulation of a PumasModel is performed via the simobs function. The function is given by the values:
Pumas.simobs — Function
simobs(
model::AbstractPumasModel,
population::Union{Subject, Population},
param,
randeffs=sample_randeffs(rng, model, population, param);
obstimes=nothing,
ensemblealg=EnsembleSerial(),
diffeq_options=NamedTuple(),
rng=Random.default_rng(),
simulate_error=Val(true),
)Simulate random observations from model for population with parameters param at obstimes (by default, use the times of the existing observations for each subject). If no randeffs is provided, then random ones are generated according to the distribution in the model.
Arguments
modelmay either be aPumasModelor aPumasEMModel.populationmay either be aPopulationofSubjects or a singleSubject.paramshould be either a single parameter set, in the form of aNamedTuple, or a vector of such parameter sets. If a vector then each of the parameter sets in that vector will be applied in turn. Example:(; tvCL=1., tvKa=2., tvV=1.)randeffsis an optional argument that, if used, should specify the random effects for each subject inpopulation. Such random effects are specified byNamedTuples forPumasModels(e.g.(; tvCL=1., tvKa=2.)) and byTuplesforPumasEMModels (e.g.(1., 2.)). Ifpopulationis a singleSubject(without being enclosed in a vector) then a single such random effect specifier should be passed. If, however,populationis aPopulationof multipleSubjects thenrandeffsshould be a vector containing one such specifier for eachSubject. The functionscenter_randeffs,zero_randeffs, andsample_randeffsare sometimes convenient for generatingrandeffs:
randeffs = zero_randeffs(model, population, param)
solve(model, population, param, randeffs)If no randeffs is provided, then random ones are generated according to the distribution in the model.
obstimesis a keyword argument where you can pass a vector of times at which to simulate observations. The default,nothing, ensures the use of the existing observation times for eachSubject.ensemblealgis a keyword argument that allows you to choose between different modes of parallelization. Options includeEnsembleSerial(),EnsembleThreads()andEnsembleDistributed().diffeq_optionsis a keyword argument that takes aNamedTupleof options to pass on to the differential equation solver. The absolute tolerance, relative tolerance, and the maximum number of iterations of the numerical solvers for computing steady states can be specified with thess_abstol,ss_reltol, andss_maxitersoptions. By default,ss_abstolandss_reltolare set to the absolute and relative tolerances of the differential equation solver, andss_maxitersis set to 1000.rngis a keyword argument where you can specify which random number generator to use.simulate_erroris a keyword argument that allows you to choose whether a sample (Val(true)ortrue) or the mean (Val(false)orfalse) of the predictive distribution's error model will be returned for each subject.
simobs(model::PumasModel, population::Population; samples::Int = 10, simulate_error = true, rng::AbstractRNG = default_rng())Simulates model predictions for population using the prior distributions for the population and subject-specific parameters. samples is the number of simulated predictions returned per subject. If simulate_error is false, the mean of the predictive distribution's error model will be returned, otherwise a sample from the predictive distribution's error model will be returned for each subject. rng is the pseudo-random number generator used in the sampling.
simobs(model::PumasModel, subject::Subject; samples::Int = 10, simulate_error = true, rng::AbstractRNG = default_rng())Simulates model predictions for subject using the prior distributions for the population and subject-specific parameters. samples is the number of simulated predictions returned. If simulate_error is false, the mean of the predictive distribution's error model will be returned, otherwise a sample from the predictive distribution's error model will be returned for each subject. rng is the pseudo-random number generator used in the sampling.
simobs(pmi::FittedPumasModelInference, population::Population = pmi.fpm.data, randeffs::Union{Nothing, AbstractVector{<:NamedTuple}} = nothing; samples::Int, rng = default_rng(), kwargs...,)Simulates observations from the fitted model using a truncated multivariate normal distribution for the parameter values when a covariance estimate is available and using non-parametric sampling for simulated inference and bootstrapping. When a covariance estimate is available, the optimal parameter values are used as the mean and the variance-covariance estimate is used as the covariance matrix. Rejection sampling is used to avoid parameter values outside the parameter domains. Each sample uses a different parameter value. samples is the number of samples to sample. population is the population of subjects which defaults to the population associated the fitted model. randeffs can be set to a vector of named tuples, one for each sample. If randeffs is not specified (the default behaviour), it will be sampled from its distribution.
simobs(fpm::FittedPumasModel, [population::Population,] vcov::AbstractMatrix, randeffs::Union{Nothing, AbstractVector{<:NamedTuple}} = nothing; samples::Int, rng = default_rng(), kwargs...)Simulates observations from the fitted model using a truncated multi-variate normal distribution for the parameter values. The optimal parameter values are used for the mean and the user supplied variance-covariance (vcov) is used as the covariance matrix. Rejection sampling is used to avoid parameter values outside the parameter domains. Each sample uses a different parameter value. samples is the number of samples to sample. population is the population of subjects which defaults to the population associated the fitted model so it's optional to pass. randeffs can be set to a vector of named tuples, one for each sample. If randeffs is not specified (the default behaviour), it will be sampled from its distribution.
For the examples showcase here, consider the following, model, population and parameters that we will use to simulate from:
using Pumas
using PumasUtilities
onecomp = @model begin
@param begin
tvcl ∈ RealDomain(; lower = 0, init = 4)
tvvc ∈ RealDomain(; lower = 0, init = 70)
Ω ∈ PDiagDomain(init = [0.04, 0.04])
σ ∈ RealDomain(; lower = 0.00001, init = 0.1)
end
@random begin
η ~ MvNormal(Ω)
end
@pre begin
CL = tvcl * exp(η[1])
Vc = tvvc * exp(η[2])
end
@dynamics Central1
@derived begin
cp := @. Central / Vc
dv ~ @. ProportionalNormal(cp, σ)
end
end
model_params = init_params(onecomp)
ext_param = (; tvcl = 3, tvvc = 60, Ω = Diagonal([0.04, 0.04]), σ = 0.1)
regimen = DosageRegimen(100)
pop = map(i -> Subject(; id = i, events = regimen, observations = (:dv,)), 1:3)
sims = simobs(onecomp, pop, ext_param; obstimes = 1:24)Simulated population (Vector{<:Subject})
Simulated subjects: 3
Simulated variables: dvOptions and settings for simobs
Below we will go through visualization and post-processing of results and some common simulation patterns in terms of subject or population level simulations and how to set or draw fixed effects and random effects. Before we get to that, let us establish how to control parallelization and settings for the ordinary differential equation (ODE) solver. The main options are:
obstimesensemblealgdiffeq_optionsrngsimulate_error
Adding times to the simulations with obstimes
When defining the subjects to be used in simulations it is possible to input the time information. This can be the values in the time column of a DataFrame with read_pumas or given directly to a subject constructor Subject(; id = "1", time = [0.0, 1.0, 5.0]). If it is desired at the time of simulation to add more time points, these can be added to the simulations using the obstimes keyword.
simobs(model, population, parameters; obstimes = [2.0, 8.0, 24.0])This will add the time points 2, 8, and 24 to the simulated time points on top of the existing time points in the subejct.
Choosing mode of parallelization ensemblealg
Simulations from simobs are multithreaded by default. This ensures that all simulations uses all the available cores or virtual CPUs on the machine being used. Other possibilities are EnsembleSerial() or EnsembleDistributed(). EnsembleSerial() is mostly present as an option for backwards compatibility. Prior to v2.6.0 of Pumas a simobs call with EnsembleThreads would not necessarily produce stable results even if a seed was set. simobs is stable across runs even with EnsembleThreads so there is no longer a good reason to use EnsembleSerial(). EnsembleDistributed() uses the distributed computing functionality in Julia. This generally means that work is divided to workers using the pmap function. Simulations are rarely sufficiently expensive per subject that this mode of parallelization makes much sense. It is possible to use it, but it is mostly relevant for large datasets with expensive model solutions in a fit context or in covariate model selection using covariate_select that always uses a pmap.
Setting numerical ODE solver options with diffeq_options
When specifying a model we very often have to include a dynamical model to account for the pharmacokinetics and pharmacodynamics. Such a model can have a closed form, or analytical, solution if it is sufficiently simple. However, in many cases there is no closed form solution for example if there are non-linear terms in the dynamics or if the model depends explicitly on t. Then, we need to solve the model numerically. This is called numerical integration because the solution of the model is the integral of the differential equations over time.
If the model is to be solved numerically there are three settings we may have to set in some circumstances. Sometimes a model does not solve well or fast enough and it is sometimes possible to change the numerical integration method (the "solver" or "algorithm") and sometimes we may want to adjust tolerances if it is impractical to solve the model to the usual tolerances. The three options we want to highlight are:
abstolroughly interpreted as the error around0reltolroughly interpreted as controlling the number of correct digits (1e-3would then mean3)algis the solver that will actually perform the numerical integration
The tolerances for the solution control the following scaled error
err_scaled = err/(abstol + (max(uprev, u)*reltol))where u and uprev are two candidates for the solution. The scaled error is guaranteed to be less than 1 and the interpretation of the two tolerances are as explained in the list above.
In Pumas, we use the AutoVern7 automatic switching algorithm with Rodas5P inside. This means that Vern7 is used by default and if stiffness is detected we switch to Rodas5P. In terms of tolerances, we set abstol=1e-12 and reltol=1e-8.
Understanding the diffeq_options in terms of NONMEM terminology
To map the advice above to what users might know from the NONMEM $SUBROUTINES functionality consider the following list:
TOLsets the relative tolerance and can be a number or a user providedTOLroutine. If the input is a numbernit must be a an integer and sets the approximate number of correct digits. In terms of Pumas options, this corresponds todiffeq_options = (; reltol = 1e-n).ATOLsets the absolute tolerance and should be an integern. This corresponds todiffeq_options = (; abstol = 1e-n)in Pumas.ADVANs such asADVAN6,ADVAN9,ADVAN13corresponds to setting thealgoption as described above. Note, that the use ofADVANdoes not exactly correspond todiffeq_options = (; alg = ...)becausealgis only use for numerical integrator choice in Pumas whereas it is also used to specify closed form solutions in NONMEM.
Turning simulation of derived distributions off or on
When running simobs there are several levels of random draws that might be made. First, if the random effects are not passed in after the fixed effects the random effects will be sampled. Second, in the @derived block all dependent variables will also be drawn. Consider the following:
@derived begin
cp = @. 1_000 * (Central / Vc)
dv1 ~ @. ProportionalNormal(cp, σ_prop)
dv2 ~ @. Normal(cp, σ_add)
endThis model would simulate both a proportional and an additive dependent variable for each subject with the same mean. If we run simobs as usual (not options set) the output will contain two variables with the same mean, but with different variances. This source of randomness is often called Residual Unexplained Variability or RUV. To run simulations with uncertainty using the parameter uncertainty of estimated fixed effects or some other simulation that requires you to simulate without RUV we use simulate_error = false
simobs(model, population, parameters, randeffs; simulate_error = false)This will use the mean of the derived variables instead of sampling from them and in this case that would mean that all dv1 and dv2 observations would be simulated to be the exact same values.
Seeding the simulations with rng
As mentioned above, simobs might sample both Between Subject Variability (BSV) or Residual Unexplained Variability (RUV). The exact values that are sampled is controlled through the random number generator (RNG).
To set a seed for the default random number generator it is possible to use code as the following:
using Random
Random.seed!(859)This will set the seed of the default RNG and as such we can simply call simobs after without no additional settings. It is also possible to assign a RNG instance to a variables and manually seed the simobs called for example using Random again:
using Random
my_rng = Random.seed!(999)
simobs(model, population, parameters; rng = my_rng)The seed! approach without manually setting the keywords should work fine in most cases though.
Seeded random sequences and simulation of sampled variables in @derived
There is an important point to make about how the random components are sampled in simobs and what it means for situations where you might want to first run a simulation for some time interval and later expand that time interval. An example could be an adaptive dosing trial where you want to first simulate up until some time point that designates a time where the dose may be modified. You look at the values, add some doses to the subjects to match their doses from the time of the visit to the clinic, and then you simulate onwards in time.
When running simobs we take the model, subject(s), fixed effects, random effects, and potentially additional obstimes and use them to first solve the underlying model. This model solution can then be used to define variables in @derived that have measurement error or some other RUV component added on top of the variables in the dynamical system as above:
@derived begin
cp = @. 1_000 * (Central / Vc)
dv1 ~ @. ProportionalNormal(cp, σ_prop)
dv2 ~ @. Normal(cp, σ_add)
endConsider the case where we are simulating the model that contains the above @derived block. If we simulate at the time points [0.0,1.0,2.0] we will then get two sequences of numbers for dv1 and dv2. These values will be sampled according to the seed that we set. If no seed is explicitly set the current global RNG state is used.
Let us imagine that we looked at the values and want to continue the simulation onwards such that the full time vector becomes [0.0,1.0,2.0,4.0,5.0,6.0] then we have to be careful in how we actually sample the @derived variables. Internally we sample all variables together per time point. First, we sample dv1 and dv2 at time = 0.0, then we sample dv1 and dv2 at time = 1.0, and so on until the final time point. This ensures that if we expand our time vector beyond the original time vector, we still get the same results out for the first part of the time vector given the same seed.
The point may be a little subtle, but it can be illustrated as follows. First, consider setting a seed and drawing two uniform values that will stand in for the dependent variables:
julia> Random.seed!(234)
TaskLocalRNG()
julia> a = rand(2)
2-element Vector{Float64}:
0.6393022684352254
0.04151237262182306
julia> b = rand(2)
2-element Vector{Float64}:
0.6035818204037028
0.8866568936656358Imagine, that we now want to simulate three values in a and b instead. Then we could re-set the seed and get:
julia> Random.seed!(234)
TaskLocalRNG()
julia> a = rand(3)
3-element Vector{Float64}:
0.6393022684352254
0.04151237262182306
0.6035818204037028
julia> b = rand(3)
3-element Vector{Float64}:
0.8866568936656358
0.952245838494935
0.7773247257534464However, this is not what we wanted! We wanted to original values from a and b and then an additional sample for each of them. If you look closely, you will see that the first three values of the new a are exactly the first two values of the original a and then first value of the original b. For b we then get the last value of the original b and then two new values.
To ensure that Pumas models always simulate the same way in the first part of the time domain even if we expand it with additional times we take another approach than sampling the full dv vectors at once. Instead, we do something analogous to the following:
julia> Random.seed!(234)
TaskLocalRNG()
julia> a_alt = []
Any[]
julia> b_alt = []
Any[]
julia> for i = 1:2
push!(a_alt, rand())
push!(b_alt, rand())
end
julia> a_alt
2-element Vector{Any}:
0.6393022684352254
0.6035818204037028
julia> b_alt
2-element Vector{Any}:
0.04151237262182306
0.8866568936656358
julia> Random.seed!(234)
TaskLocalRNG()
julia> a_alt = []
Any[]
julia> b_alt = []
Any[]
julia> for i = 1:3
push!(a_alt, rand())
push!(b_alt, rand())
end
julia> a_alt
3-element Vector{Any}:
0.6393022684352254
0.6035818204037028
0.952245838494935
julia> b_alt
3-element Vector{Any}:
0.04151237262182306
0.8866568936656358
0.7773247257534464While it is beyond the scope of this document to show a full example this is important for the use-case mentioned above where you have an adaptive trial and you want to sequentially run the simobs up until some time point, check the state of the subjects, add some doses to the subject, and then simobs again but this time going further to the next visit to the clinic.
Visualizing Simulated Returns
The object returned by simobs have automatic plotting and DataFrame conversion. To plot a simulation return, simply call sim_plot on the output:
sim_plot(sims)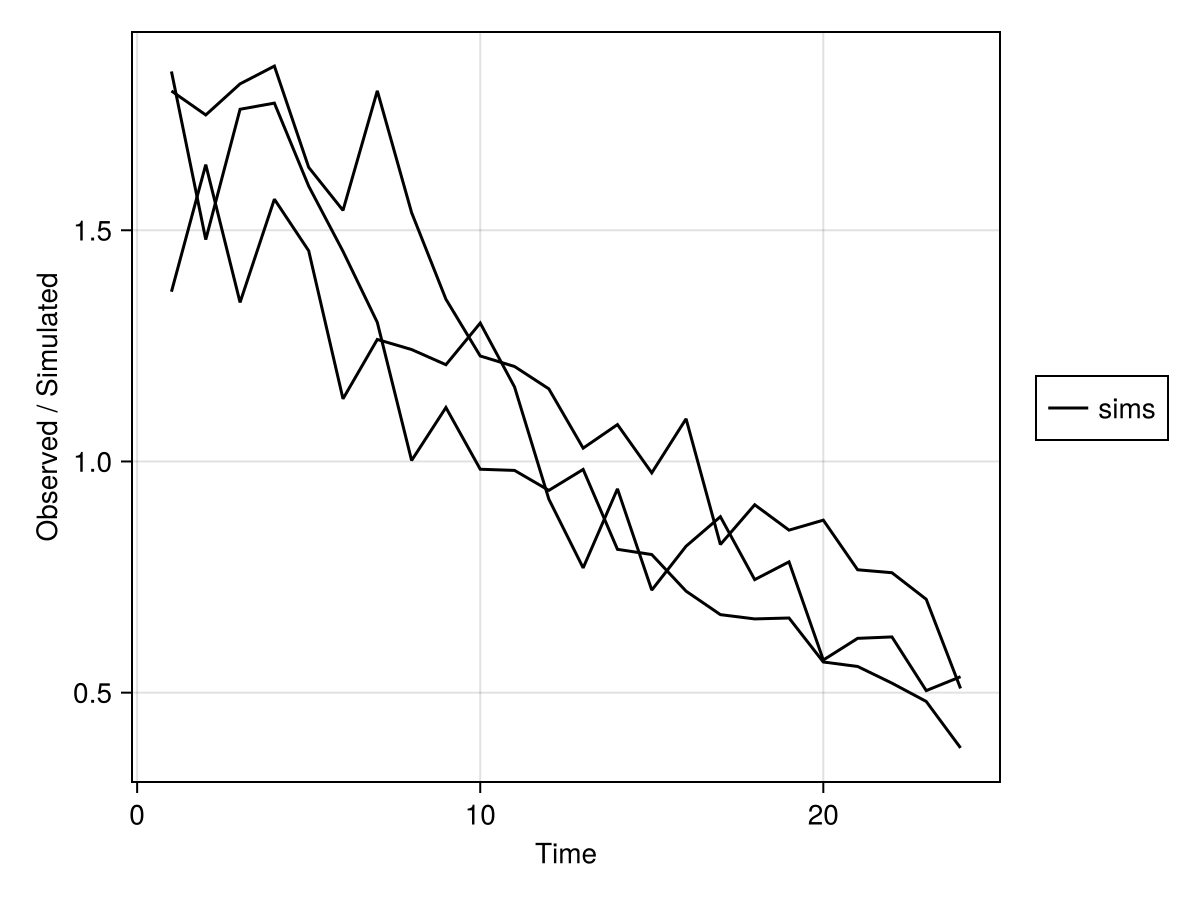
This generates a plot for each variable(s) specified in observations keyword. For example, sim_plot(m, obs; observations=[:dv1, :dv2]) would only plot the values dv1 and dv2. In addition, all the specified attributes can be used in this plot command. For more advanced use of the underlying plotting ecosystem, please see the Makie layout tutorial. Note that if the simulated return is a SimulatedPopulation, then the plots overlay the results of the various subjects.
To generate the DataFrame associated with the observed outputs, simply call DataFrame on the simulated return. For example, the following builds the tabular output from the returned object:
df = DataFrame(sims)
first(df, 5)| Row | id | time | dv | evid | amt | cmt | rate | duration | ss | ii | route | tad | dosenum | Central | η₁ | η₂ | CL | Vc |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| String | Float64 | Float64? | Int64? | Float64? | Symbol? | Float64? | Float64? | Int8? | Float64? | NCA.Route? | Float64? | Int64? | Float64? | Float64? | Float64? | Float64? | Float64? | |
| 1 | 1 | 0.0 | missing | 1 | 100.0 | Central | 0.0 | 0.0 | 0 | 0.0 | NullRoute | 0.0 | 1 | missing | -0.243451 | -0.234909 | 2.35175 | 47.4386 |
| 2 | 1 | 1.0 | 1.80126 | 0 | 0.0 | missing | 0.0 | 0.0 | 0 | 0.0 | missing | 1.0 | 1 | 95.1634 | -0.243451 | -0.234909 | 2.35175 | 47.4386 |
| 3 | 1 | 2.0 | 1.74937 | 0 | 0.0 | missing | 0.0 | 0.0 | 0 | 0.0 | missing | 2.0 | 1 | 90.5607 | -0.243451 | -0.234909 | 2.35175 | 47.4386 |
| 4 | 1 | 3.0 | 1.81664 | 0 | 0.0 | missing | 0.0 | 0.0 | 0 | 0.0 | missing | 3.0 | 1 | 86.1807 | -0.243451 | -0.234909 | 2.35175 | 47.4386 |
| 5 | 1 | 4.0 | 1.85505 | 0 | 0.0 | missing | 0.0 | 0.0 | 0 | 0.0 | missing | 4.0 | 1 | 82.0125 | -0.243451 | -0.234909 | 2.35175 | 47.4386 |
Different usage patterns of simobs
Below, we showcase the different usage patterns of simobs.
Subject level simulation
In this pattern, you can use simobs to do a single subject simulation by passing a Subject instead of a Population:
sims2 = simobs(onecomp, pop[1], ext_param; obstimes = 1:24)SimulatedObservations
Simulated variables: dv
Time: 1:24where, onecomp is the model, pop[1] represents one subject in the Population, pop. The parameters of the model are passed in as the NamedTuple defined earlier.
Subject level simulation with passing random effects
In this pattern, you can use simobs to do a single subject simulation by passing an additional positional argument of individual subject random effects represented as vector. The length of the random effect vector should be the same as the number of parameters for each subject:
ext_randeffs = (; η = [0.7, -0.44])
sims3 = simobs(onecomp, pop[1], ext_param, ext_randeffs; obstimes = 1:24)SimulatedObservations
Simulated variables: dv
Time: 1:24where, onecomp is the model, pop[1] represents one subject in the Population, pop. The parameters of the model are passed in as the NamedTuple defined earlier. ext_randeffs are the specific values of the η's that are passed in, which means that simobs uses these values as opposed to those sampled from the distributions specified in the @random block.
Subject level simulation with passing in an array of parameters
In this pattern, you can use simobs to do a single subject simulation by passing a positional argument, an array of parameters that generates a unique simulation solution for each element of the parameter array. E.g. let's simulate pop[1] with three different combinations of tvcl and tvvc. In the example below, we change the value of tvcl to represent lo, med, hi of 3, 6, 9, but we don't change the other values (Note that are more efficient programmatic ways to generate such arrays) using the syntax below:
ext_param_array = [
(tvcl = 3, tvvc = 60, Ω = Diagonal([0.04, 0.04]), σ = 0.1),
(tvcl = 6, tvvc = 60, Ω = Diagonal([0.04, 0.04]), σ = 0.1),
(tvcl = 9, tvvc = 60, Ω = Diagonal([0.04, 0.04]), σ = 0.1),
]
sims4 = simobs(onecomp, pop[1], ext_param_array; obstimes = 1:24)Simulated population (Vector{<:Subject})
Simulated subjects: 3
Simulated variables: dvwhere, onecomp is the model, pop[1] represents one subject in the Population, pop. The parameters of the model are passed in as the vector of NamedTuples ext_param_array.
A DataFrame of parameter values can be converted to a NamedTuple easily using Tables.rowtable syntax. See the example below.
myparams = DataFrame(;
tvcl = [1, 2, 3],
tvvc = [20, 30, 40],
Ω = [Diagonal([0.04, 0.04]), Diagonal([0.04, 0.04]), Diagonal([0.04, 0.04])],
σ = 0.1,
)| Row | tvcl | tvvc | Ω | σ |
|---|---|---|---|---|
| Int64 | Int64 | Diagonal… | Float64 | |
| 1 | 1 | 20 | [0.04 0.0; 0.0 0.04] | 0.1 |
| 2 | 2 | 30 | [0.04 0.0; 0.0 0.04] | 0.1 |
| 3 | 3 | 40 | [0.04 0.0; 0.0 0.04] | 0.1 |
This can be converted to a NamedTuple
myparam_tuple = Tables.rowtable(myparams)3-element Vector{@NamedTuple{tvcl::Int64, tvvc::Int64, Ω::Diagonal{Float64, Vector{Float64}}, σ::Float64}}:
(tvcl = 1, tvvc = 20, Ω = [0.04 0.0; 0.0 0.04], σ = 0.1)
(tvcl = 2, tvvc = 30, Ω = [0.04 0.0; 0.0 0.04], σ = 0.1)
(tvcl = 3, tvvc = 40, Ω = [0.04 0.0; 0.0 0.04], σ = 0.1)myparam_tuple is of the same form as the ext_param_array.
Subject level simulation with passing in an array of parameters and random effects
The two usage patterns above can be combined such that one can pass in an array of parameters and corresponding random effects with the usage pattern below:
ext_randeffs = [(; η = [0.7, -0.44]), (; η = [0.6, -0.9]), (; η = [-0.7, 0.6])]
sims5 = simobs(onecomp, pop[1], ext_param_array, ext_randeffs; obstimes = 1:24)Simulated population (Vector{<:Subject})
Simulated subjects: 3
Simulated variables: dvPopulation level simulations
In this pattern, you can use simobs to do a whole population simulation by passing a Population instead of a Subject:
sims6 = simobs(onecomp, pop, ext_param; obstimes = 1:24)Simulated population (Vector{<:Subject})
Simulated subjects: 3
Simulated variables: dvwhere, onecomp is the model, pop is the Population. The parameters of the model are passed in as the NamedTuple defined earlier.
Population level simulation passing random effects
In this pattern, you can use simobs to do a population simulation by passing an additional positional argument of individual subject random effects represented as vector. The length of the random effect vector should be the same as the number of parameters for each subject:
You can use this method to pass in the empirical_bayes of a FittedPumasModel.
ext_randeffs = [(; η = rand(2)), (; η = rand(2)), (; η = rand(2))]
sims8 = simobs(onecomp, pop, ext_param, ext_randeffs; obstimes = 1:24)Simulated population (Vector{<:Subject})
Simulated subjects: 3
Simulated variables: dvwhere, onecomp is the model, pop represents the Population, pop. The parameters of the model are explicitly passed in as the NamedTuple defined earlier. ext_randeffs are the specific values of the η's that are passed in, which means that simobs uses these values as opposed to those sampled from the distributions specified in the @random block.
Population level simulation with passing in an array of parameters
The two usage patterns above can be combined such that one can pass in an array of parameters and corresponding random effects.
Here, a population can be simulated with an array of parameters that generates a unique simulation solution for each element of the parameter array. E.g. let's simulate pop with three different combinations of tvcl and tvvc. In the example below, we change the value of tvcl to represent lo, med, hi of 3, 6, 9, but we don't change the other values (Note that are more efficient programmatic ways to generate such arrays):
ext_param_array = [
(; tvcl = 3, tvvc = 60, Ω = Diagonal([0.04, 0.04]), σ = 0.1),
(; tvcl = 6, tvvc = 60, Ω = Diagonal([0.04, 0.04]), σ = 0.1),
(; tvcl = 9, tvvc = 60, Ω = Diagonal([0.04, 0.04]), σ = 0.1),
]
sims9 = simobs(onecomp, pop, ext_param_array; obstimes = 1:24)3-element Vector{Vector{Pumas.SimulatedObservations{PumasModel{(tvcl = 1, tvvc = 1, Ω = 2, σ = 1), 2, (:Central,), (:CL, :Vc), ParamSet{@NamedTuple{tvcl::RealDomain{Int64, TransformVariables.Infinity{true}, Int64}, tvvc::RealDomain{Int64, TransformVariables.Infinity{true}, Int64}, Ω::PDiagDomain{PDMats.PDiagMat{Float64, Vector{Float64}}}, σ::RealDomain{Float64, TransformVariables.Infinity{true}, Float64}}}, ParamSet{@NamedTuple{η::Pumas.RandomEffect{false, Main.var"#1#6"}}}, Pumas.PreObj{Main.var"#2#7"}, Pumas.DCPObj{Returns{Returns{@NamedTuple{}}}}, Main.var"#3#8", Central1, Pumas.DerivedObj{(:dv,), false, Main.var"#4#9"}, Main.var"#5#10", Nothing, Pumas.PumasModelOptions}, Subject{@NamedTuple{dv::Vector{Missing}}, Pumas.ConstantCovar{@NamedTuple{}}, DosageRegimen{Float64, Symbol, Float64, Float64, Float64, Float64}, Nothing}, UnitRange{Int64}, @NamedTuple{tvcl::Int64, tvvc::Int64, Ω::Diagonal{Float64, Vector{Float64}}, σ::Float64}, @NamedTuple{η::Vector{Float64}}, @NamedTuple{}, @NamedTuple{saveat::UnitRange{Int64}, ss_abstol::Float64, ss_reltol::Float64, ss_maxiters::Int64, abstol::Float64, reltol::Float64, alg::CompositeAlgorithm{0, Tuple{Vern7{typeof(OrdinaryDiffEqCore.trivial_limiter!), typeof(OrdinaryDiffEqCore.trivial_limiter!), Static.False}, Rodas5P{1, ADTypes.AutoForwardDiff{1, Nothing}, LinearSolve.GenericLUFactorization{RowMaximum}, typeof(OrdinaryDiffEqCore.DEFAULT_PRECS), Val{:forward}(), true, nothing, typeof(OrdinaryDiffEqCore.trivial_limiter!), typeof(OrdinaryDiffEqCore.trivial_limiter!)}}, OrdinaryDiffEqCore.AutoSwitch{Vern7{typeof(OrdinaryDiffEqCore.trivial_limiter!), typeof(OrdinaryDiffEqCore.trivial_limiter!), Static.False}, Rodas5P{1, ADTypes.AutoForwardDiff{1, Nothing}, LinearSolve.GenericLUFactorization{RowMaximum}, typeof(OrdinaryDiffEqCore.DEFAULT_PRECS), Val{:forward}(), true, nothing, typeof(OrdinaryDiffEqCore.trivial_limiter!), typeof(OrdinaryDiffEqCore.trivial_limiter!)}, Rational{Int64}, Int64}}, continuity::Symbol}, Pumas.PKPDAnalyticalSolution{StaticArraysCore.SVector{1, Float64}, 2, Vector{StaticArraysCore.SVector{1, Float64}}, Vector{Float64}, Vector{StaticArraysCore.SVector{1, Float64}}, Vector{StaticArraysCore.SVector{1, Float64}}, Returns{@NamedTuple{CL::Float64, Vc::Float64}}, Pumas.AnalyticalPKPDProblem{StaticArraysCore.SVector{1, Float64}, Float64, false, Central1, Vector{Pumas.Event{Float64, Float64, Float64}}, Vector{Float64}, Returns{@NamedTuple{CL::Float64, Vc::Float64}}}}, @NamedTuple{}, @NamedTuple{CL::Vector{Float64}, Vc::Vector{Float64}}, @NamedTuple{dv::Vector{Float64}}, @NamedTuple{}}}}:
Simulated population (Vector{<:Subject}), n = 3, variables: dv
Simulated population (Vector{<:Subject}), n = 3, variables: dv
Simulated population (Vector{<:Subject}), n = 3, variables: dvwhere, onecomp is the model, pop represents the Population, pop. The parameters of the model are passed in as a vector of the NamedTuples ext_param_array as defined earlier represents the array of parameter values we want to simulate the model at.
Population level simulation with passing in an array of parameters and random effects
The two usage patterns above can be combined such that one can pass in an array of parameters and corresponding random effects with the usage pattern below:
ext_randeffs_array = [(; η = rand(2)) for i = 1:length(ext_param_array)*length(pop)]
sims10 = simobs(onecomp, pop, ext_param_array, ext_randeffs_array; obstimes = 1:24)3-element Vector{Vector{Pumas.SimulatedObservations{PumasModel{(tvcl = 1, tvvc = 1, Ω = 2, σ = 1), 2, (:Central,), (:CL, :Vc), ParamSet{@NamedTuple{tvcl::RealDomain{Int64, TransformVariables.Infinity{true}, Int64}, tvvc::RealDomain{Int64, TransformVariables.Infinity{true}, Int64}, Ω::PDiagDomain{PDMats.PDiagMat{Float64, Vector{Float64}}}, σ::RealDomain{Float64, TransformVariables.Infinity{true}, Float64}}}, ParamSet{@NamedTuple{η::Pumas.RandomEffect{false, Main.var"#1#6"}}}, Pumas.PreObj{Main.var"#2#7"}, Pumas.DCPObj{Returns{Returns{@NamedTuple{}}}}, Main.var"#3#8", Central1, Pumas.DerivedObj{(:dv,), false, Main.var"#4#9"}, Main.var"#5#10", Nothing, Pumas.PumasModelOptions}, Subject{@NamedTuple{dv::Vector{Missing}}, Pumas.ConstantCovar{@NamedTuple{}}, DosageRegimen{Float64, Symbol, Float64, Float64, Float64, Float64}, Nothing}, UnitRange{Int64}, @NamedTuple{tvcl::Int64, tvvc::Int64, Ω::Diagonal{Float64, Vector{Float64}}, σ::Float64}, @NamedTuple{η::Vector{Float64}}, @NamedTuple{}, @NamedTuple{saveat::UnitRange{Int64}, ss_abstol::Float64, ss_reltol::Float64, ss_maxiters::Int64, abstol::Float64, reltol::Float64, alg::CompositeAlgorithm{0, Tuple{Vern7{typeof(OrdinaryDiffEqCore.trivial_limiter!), typeof(OrdinaryDiffEqCore.trivial_limiter!), Static.False}, Rodas5P{1, ADTypes.AutoForwardDiff{1, Nothing}, LinearSolve.GenericLUFactorization{RowMaximum}, typeof(OrdinaryDiffEqCore.DEFAULT_PRECS), Val{:forward}(), true, nothing, typeof(OrdinaryDiffEqCore.trivial_limiter!), typeof(OrdinaryDiffEqCore.trivial_limiter!)}}, OrdinaryDiffEqCore.AutoSwitch{Vern7{typeof(OrdinaryDiffEqCore.trivial_limiter!), typeof(OrdinaryDiffEqCore.trivial_limiter!), Static.False}, Rodas5P{1, ADTypes.AutoForwardDiff{1, Nothing}, LinearSolve.GenericLUFactorization{RowMaximum}, typeof(OrdinaryDiffEqCore.DEFAULT_PRECS), Val{:forward}(), true, nothing, typeof(OrdinaryDiffEqCore.trivial_limiter!), typeof(OrdinaryDiffEqCore.trivial_limiter!)}, Rational{Int64}, Int64}}, continuity::Symbol}, Pumas.PKPDAnalyticalSolution{StaticArraysCore.SVector{1, Float64}, 2, Vector{StaticArraysCore.SVector{1, Float64}}, Vector{Float64}, Vector{StaticArraysCore.SVector{1, Float64}}, Vector{StaticArraysCore.SVector{1, Float64}}, Returns{@NamedTuple{CL::Float64, Vc::Float64}}, Pumas.AnalyticalPKPDProblem{StaticArraysCore.SVector{1, Float64}, Float64, false, Central1, Vector{Pumas.Event{Float64, Float64, Float64}}, Vector{Float64}, Returns{@NamedTuple{CL::Float64, Vc::Float64}}}}, @NamedTuple{}, @NamedTuple{CL::Vector{Float64}, Vc::Vector{Float64}}, @NamedTuple{dv::Vector{Float64}}, @NamedTuple{}}}}:
Simulated population (Vector{<:Subject}), n = 3, variables: dv
Simulated population (Vector{<:Subject}), n = 3, variables: dv
Simulated population (Vector{<:Subject}), n = 3, variables: dvSimulation using a FittedPumasModelInference
The usage pattern is represented below:
Pumas.simobs — Method
simobs(pmi::FittedPumasModelInference, population::Population = pmi.fpm.data, randeffs::Union{Nothing, AbstractVector{<:NamedTuple}} = nothing; samples::Int, rng = default_rng(), kwargs...,)Simulates observations from the fitted model using a truncated multivariate normal distribution for the parameter values when a covariance estimate is available and using non-parametric sampling for simulated inference and bootstrapping. When a covariance estimate is available, the optimal parameter values are used as the mean and the variance-covariance estimate is used as the covariance matrix. Rejection sampling is used to avoid parameter values outside the parameter domains. Each sample uses a different parameter value. samples is the number of samples to sample. population is the population of subjects which defaults to the population associated the fitted model. randeffs can be set to a vector of named tuples, one for each sample. If randeffs is not specified (the default behaviour), it will be sampled from its distribution.
Any model that has been fitted results in a FittedPumasModelInference. simobs can simulate from the inferred object where it utilizes the variance-covariance-matrix of the infer and the design of the data that was fitted.
Post-processing simulations
The output of a simobs call stores the simulated observations but also all the intermediate values computed such as the parameters used, individual coefficients, dose control parameters, covariates, differential equation solution, etc. There are a number of post-processing operations you can do on the simulation output to compute various queries and summary statistics based on the simulations.
The postprocess function is a powerful tool that allows you to make various queries using the simulation results.
Pumas.postprocess — Function
postprocess(obs::SimulatedObservations)
postprocess(func::Function, obs::SimulatedObservations)
postprocess(obs::SimulatedPopulation; stat = identity)
postprocess(func::Function, obs::SimulatedPopulation; stat = identity)Post-processes the output of simobs. The output of a simobs call stores the simulated observations but also all the intermediate values computed such as the parameters used, individual coefficients, dose control parameters, covariates, differential equation solution, etc. There are a number of post-processing operations you can do on the simulation output to compute various queries and summary statistics based on the simulations.
The postprocess function is a powerful tool that allows you to make various queries using the simulation results. There are multiple ways to use the postprocess function. The first way to use the postprocess function is to extract all the information stored in the simulation result in the form of a vector of named tuples. Each named tuple has all the intermediate values evaluated when simulating 1 run. Let sims be the output of any simobs operation.
generated = postprocess(sims)generated is a vector of simulation runs. Hence, generated[i] is the result from the ith simulation run. Time-dependent variables are stored as a vector with 1 element for each time point where there is an observation. Alternatively, if obstimes was set instead when calling simobs, the time-dependent variables will be evaluated at the time points in obstimes instead.
The second way to use postprocess is by passing in a post-processing function. The post-processing function can be used to:
- Transform the simulated quantities, or
- Compare the simulated quantities to the observations.
We use the do syntax here which is short for passing in a function as the first argument to postprocess. The do syntax to pass in a post-processing function is:
postprocess(sims) do gen, obs
...
endwhere gen is the named tuple of all generated quantities from 1 simulation run and obs is the named tuple of observations. For instance to query the ratio of simulated observations dv that are higher than the observed quantity dv at the respective time points, you can use:
postprocess(sims) do gen, obs
sum(gen.dv .> obs.dv) / length(gen.dv)
endgen.dv is the simulated vector of dv whose length is the same as the number of observation time points. obs.dv is the observation vector dv. gen.dv .> obs.dv returns a vector of true/false, with one element for each time point. The sum of this vector gives the number of time points where the simulation was higher than the observation. Dividing by the number of time points gives the ratio. When using postprocess in this way, the output is always a vector of the query results. In the query function body, you can choose to use only gen or only obs but the header must always have both gen and obs:
postprocess(sims) do gen, obs
...
endThe third way to use the postprocess function is to compute summary statistics of the simulated quantities or of functions of the simulated quantities. Summary statistics can be computed by passing a statistic function as the stat keyword argument. For example in order to estimate the probability that a simulated value is higher than an observation, you can use:
postprocess(sims, stat = mean) do gen, obs
gen.dv .> obs.dv
endThis function will do 2 things:
- Concatenate the query results (e.g.
gen.dv .> obs.dv) from all the simulation runs into a single vector. - Compute the mean value of the combined vector.
In order to get the probability that the simulated quantity is higher than the observation for each time point, you can call the mean function externally as such:
generated = postprocess(sims) do gen, obs
gen.dv .> obs.dv
end
mean(generated)This returns a vector of probabilities of the same length as the number of time points without concatenating all the queries together.
To compute a summary statistic of all the generated quantity, you can also call:
postprocess(sims, stat = mean)without specifying a post-processing function.
Besides mean, you can also use any of the following summary statistic functions in the same way:
stdfor element-wise standard deviationvarfor element-wise variancecorfor correlation between multiple quantitiescovfor covariance between multiple quantities
These functions can be passed in as the stat keyword argument to postprocess, e.g.,
generated = postprocess(sims, stat = std) do gen, obs
...
endor
generated = postprocess(sims, stat = var) do gen, obs
...
endThe cor and cov statistics are unique in that they require a post-processing function that outputs a vector. For example, to estimate the correlation between the CL and Vc parameters in a 1-compartment model, you can use any of the following:
postprocess(s, stat = cor) do gen, obs
[only(unique(gen.CL)), only(unique(gen.Vc))]
endcov can be used instead of cor to compute the covariance matrix. The output of this operation is either a correlation or a covariance matrix.