Non-Compartmental Analysis (NCA)
NCA in Pumas is conducted in the following steps:
- Read source data (
.csv,.xlxs,.xpt,sas7bdata) - Create a
NCAPopulationby mapping variables from source data to Pumas-NCA data format - PumasNCADF - Exploratory data analysis
- Run
NCAanalysis - Generate report
Pumas also provides a convenient way to perform NCA analysis within the model as well, this is discussed in the Model integrated NCA section.
Read Source Data
The Pumas-NCA data format - PumasNCADF provides the specification requirements of the source data. Currently, several file formats can be read in for analysis:
.csv(CSVpackage).xslx(XSLXpackage).sas7bdat/.xpt(ReadStatTablespackage).arrow(Arrowpackage)
For this introduction we'll use the PharmaDatasets package and the po_sad_1 dataset:
using PharmaDatasets
pkdata = dataset("po_sad_1")
first(pkdata, 6)| Row | id | time | dv | amt | evid | cmt | rate | age | wt | doselevel | isPM | isfed | sex | route |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Int64 | Float64 | Float64? | Float64? | Int64 | Int64? | Float64 | Int64 | Int64 | Int64 | String3 | String3 | String7 | String3 | |
| 1 | 1 | 0.0 | missing | 30.0 | 1 | 1 | 0.0 | 51 | 74 | 30 | no | yes | male | ev |
| 2 | 1 | 0.25 | 35.7636 | missing | 0 | missing | 0.0 | 51 | 74 | 30 | no | yes | male | ev |
| 3 | 1 | 0.5 | 71.9551 | missing | 0 | missing | 0.0 | 51 | 74 | 30 | no | yes | male | ev |
| 4 | 1 | 0.75 | 97.3356 | missing | 0 | missing | 0.0 | 51 | 74 | 30 | no | yes | male | ev |
| 5 | 1 | 1.0 | 128.919 | missing | 0 | missing | 0.0 | 51 | 74 | 30 | no | yes | male | ev |
| 6 | 1 | 2.0 | 155.85 | missing | 0 | missing | 0.0 | 51 | 74 | 30 | no | yes | male | ev |
Create a NCAPopulation
The generated DataFrame object now needs to be mapped to the requirements of Pumas NCA package. Mapping is done using the read_nca function that is described in detail later. An example for read_nca is the following:
using NCA
ncapop = read_nca(pkdata; observations = :dv, group = [:doselevel])NCAPopulation (18 subjects):
Group: [["doselevel" => 30], ["doselevel" => 60], ["doselevel" => 90]]
Number of missing observations: 18
Number of blq observations: 0Mapping a DataFrame to the Pumas NCA package requirements using read_nca generates an object called NCAPopulation, which is collection of NCASubject's, i.e., ncapop is of type NCAPopulation.
Exploratory Data Analysis
Exploratory analysis of the NCAPopulation can be performed using the built-in plotting ecosystem. The NCA subsection of the Plotting section provides more details, but here are some example syntax and the corresponding plots.
Observations-time plots
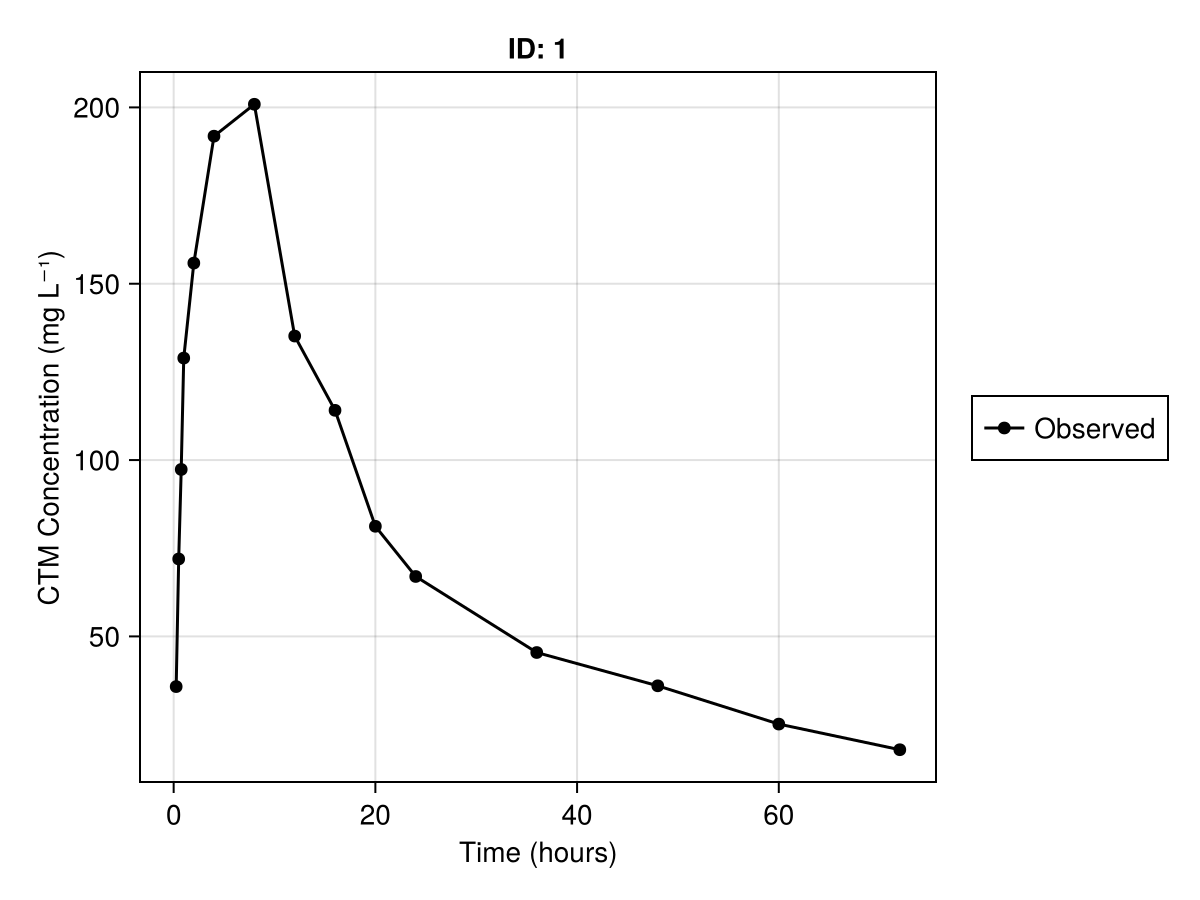
We can produce a plot of the concentration against time using observations_vs_time. The plotting recipes for NCA analyses are fund in the NCAUtilities package. Below is the code for generating this plot for the first subject found using ncapop[1]:
using NCAUtilities
observations_vs_time(
ncapop[1];
axis = (; xlabel = "Time (hours)", ylabel = "CTM Concentration (mg L⁻¹)"),
)
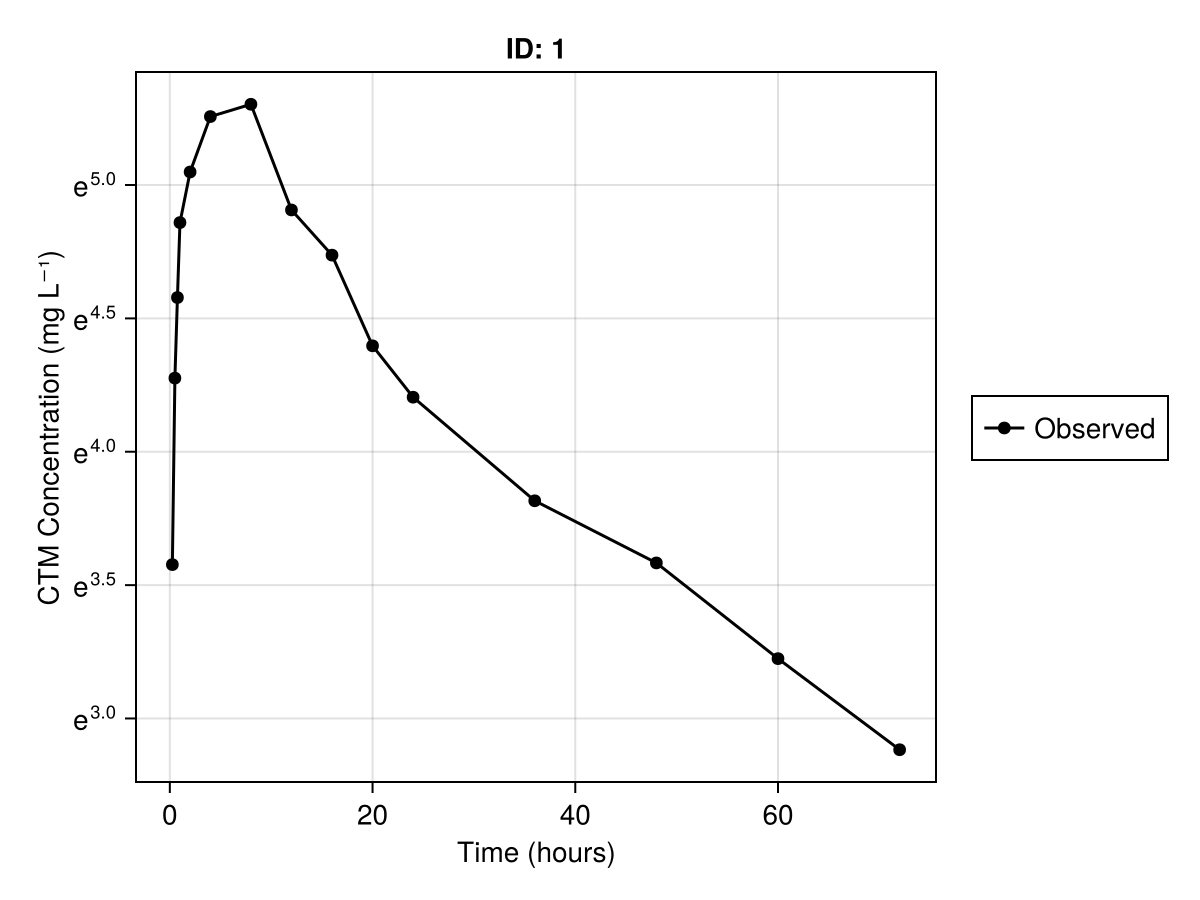
We can also use log-scale for the concentrations such that exponential decay shows as a straight line. Below is the code to set the y-scale to use the natural logarithm:
observations_vs_time(
ncapop[1];
axis = (; xlabel = "Time (hours)", ylabel = "CTM Concentration (mg L⁻¹)", yscale = log),
)
Other options are yscale = log10 and yscale = log2 for a base-10 and base-2 logarithm.
To generate these plots side-by-side we have to create a Figure-object first, and then pass fig[row,column] as the first argument. row specifies which row of the figure the element should be added to and column specifies the column number.
using CairoMakie
fig = Figure()
observations_vs_time(
fig[1, 1],
ncapop[1];
axis = (; xlabel = "Time (hours)", ylabel = "CTM Concentration (mg L⁻¹)"),
)
observations_vs_time(
fig[1, 2],
ncapop[1];
axis = (; xlabel = "Time (hours)", ylabel = "CTM Concentration (mg L⁻¹)", yscale = log),
)
fig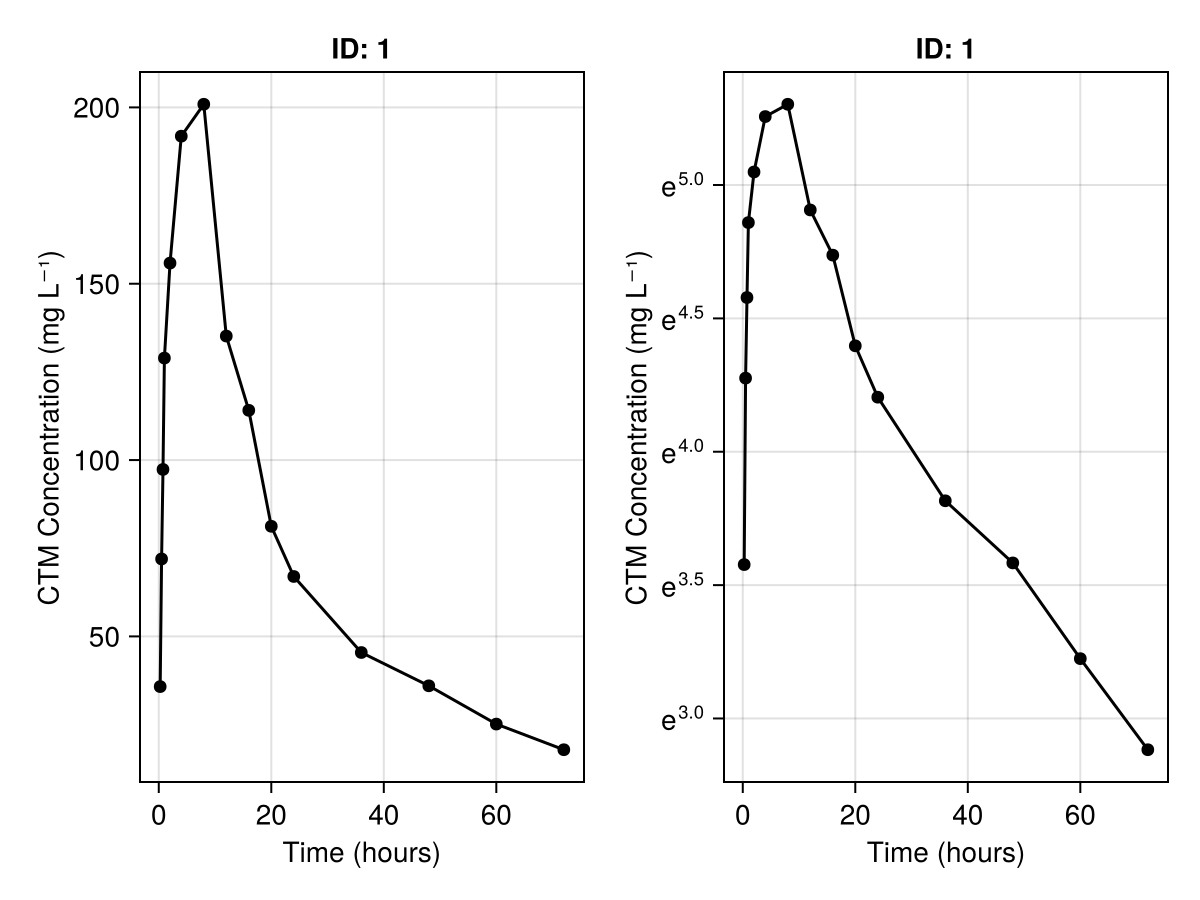
Summary plots
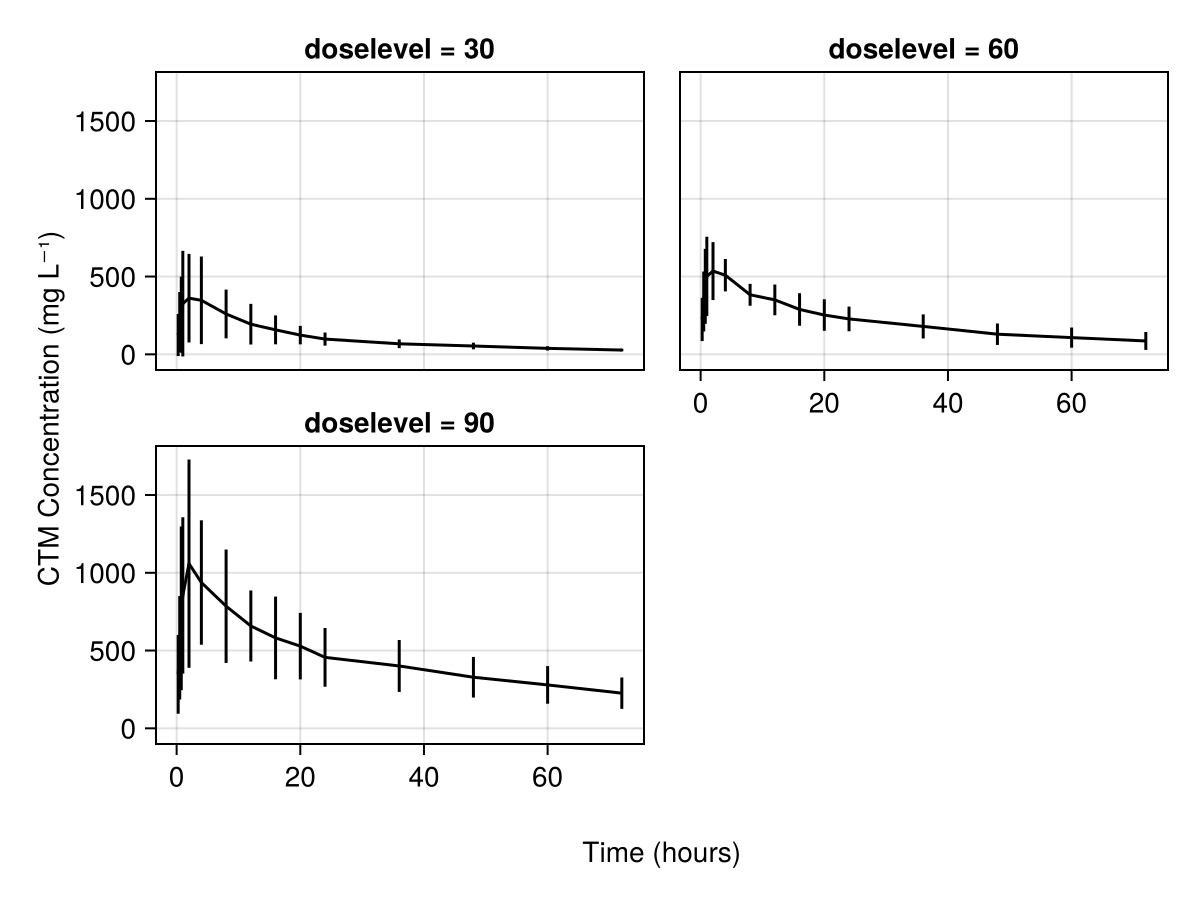
We can also group all the subjects into a single summarizing plot of the concentration against time using summary_observations_vs_time:
summary_observations_vs_time(
ncapop;
axis = (; xlabel = "Time (hours)", ylabel = "CTM Concentration (mg L⁻¹)"),
)
Subject fit plots
We can also check all the subject fits with:
subject_fits(ncapop; axis = (; yscale = log), rows = 3, columns = 3)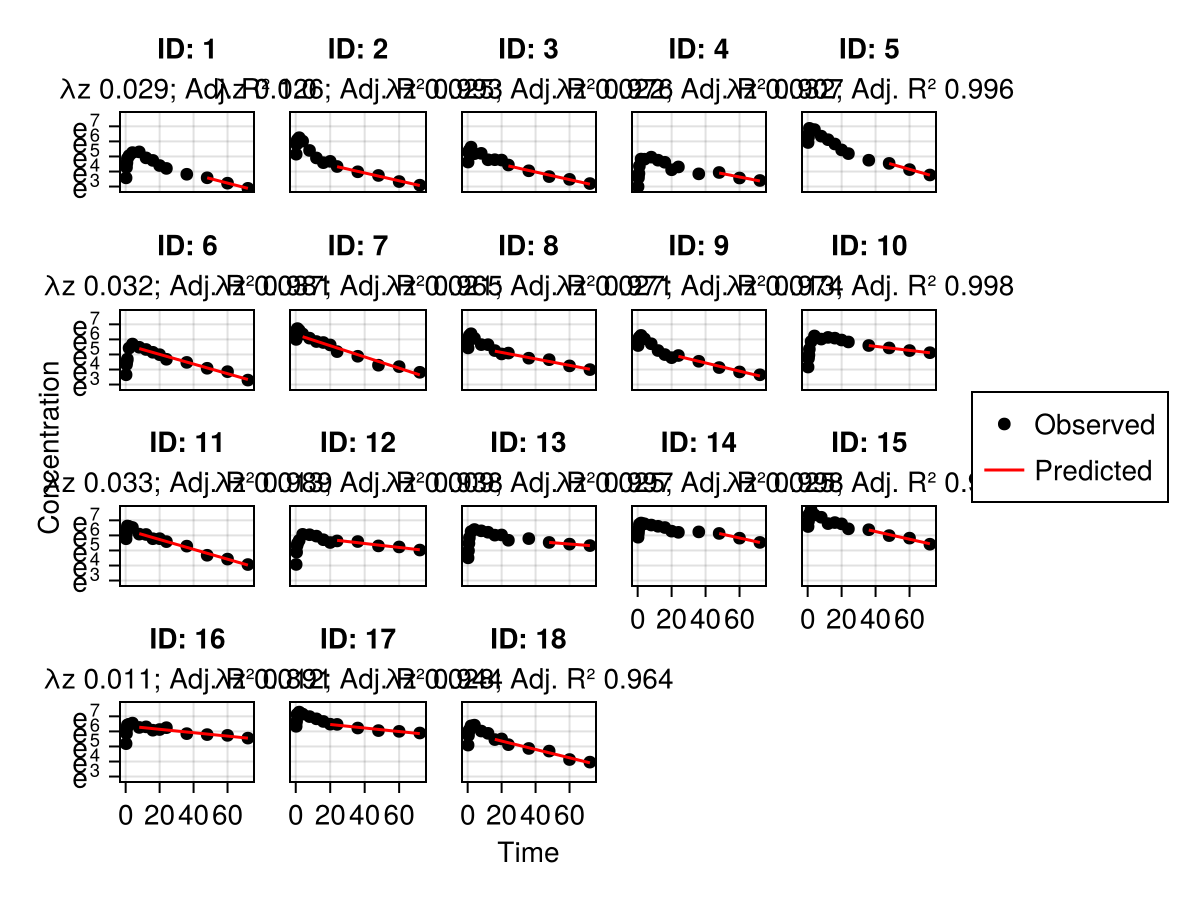
Run Analysis
The NCAPopulation object is the processed version of the DataFrame that is amenable for a complete NCA analysis via run_nca or individual parameter results through a collection of functions described in the NCA Function List. Example syntax to perform a complete NCA analysis is below:
using Dates
pk_nca = run_nca(
ncapop;
sigdigits = 3,
studyid = "STUDY-001",
studytitle = "Phase 1 SAD of Drug Y",
author = [("Mary Jane", "Pumas-AI"), ("Joe Smith", "Pumas-AI")],
sponsor = "PumasAI",
date = Dates.now(),
conclabel = "CTMX Concentration (mg/L)",
grouplabels = ["Dose (mg)"],
timelabel = "Time (Hr)",
versionnumber = v"0.1",
)The result of a complete NCA analysis using run_nca is an object called NCAReport, i.e., pk_nca is of type NCAReport. This object carries the result of the analysis in a DataFrame called reportdf and corresponding metadata information that are used for post-processing the results.
Summarize results
The per subject results in the DataFrame component of NCAReport, reportdf, can be summarized using the summarize function:
parms = [:cmax, :aucinf_obs]
summary_output = summarize(pk_nca.reportdf; parameters = parms)| Row | parameters | numsamples | minimum | maximum | mean | std | geomean | geostd | geomeanCV |
|---|---|---|---|---|---|---|---|---|---|
| String | Int64 | Float64 | Float64 | Float64 | Float64 | Float64 | Float64 | Float64 | |
| 1 | cmax | 18 | 142.0 | 2230.0 | 696.0 | 493.824 | 567.057 | 1.94997 | 74.9677 |
| 2 | aucinf_obs | 18 | 5600.0 | 75400.0 | 26070.0 | 21122.4 | 18866.5 | 2.32614 | 101.954 |
The summary_output generated above is of type DataFrame.
Plot results
We can use several plotting functions from NCAUtilities package for the results of run_nca.
The NCA subsection of the Plotting section provides more details, but here are some example syntax and the corresponding plots.
Subject fits
subject_fits(pk_nca; columns = 2, rows = 3)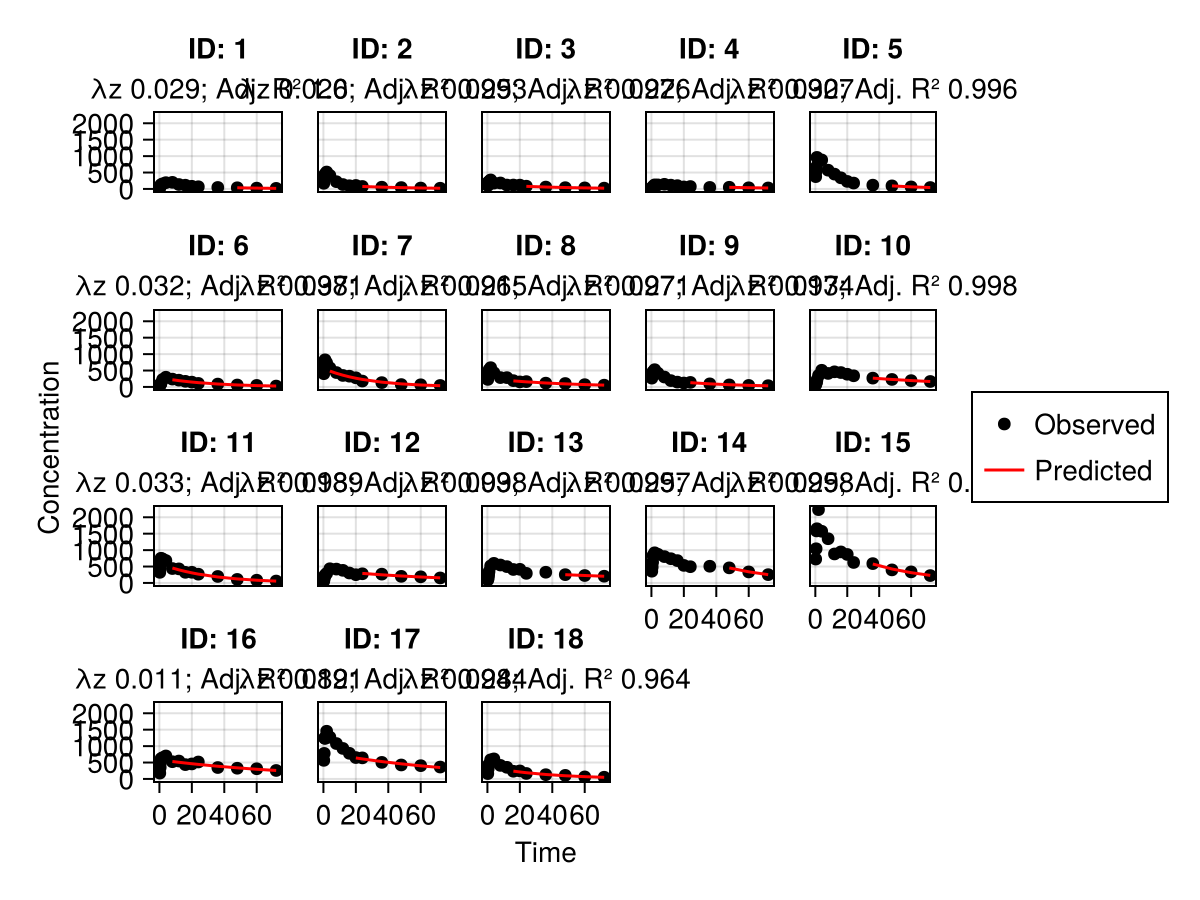
Parameter distributions
parameters_dist(pk_nca; parameter = :aucinf_obs, rows = 3, columns = 1)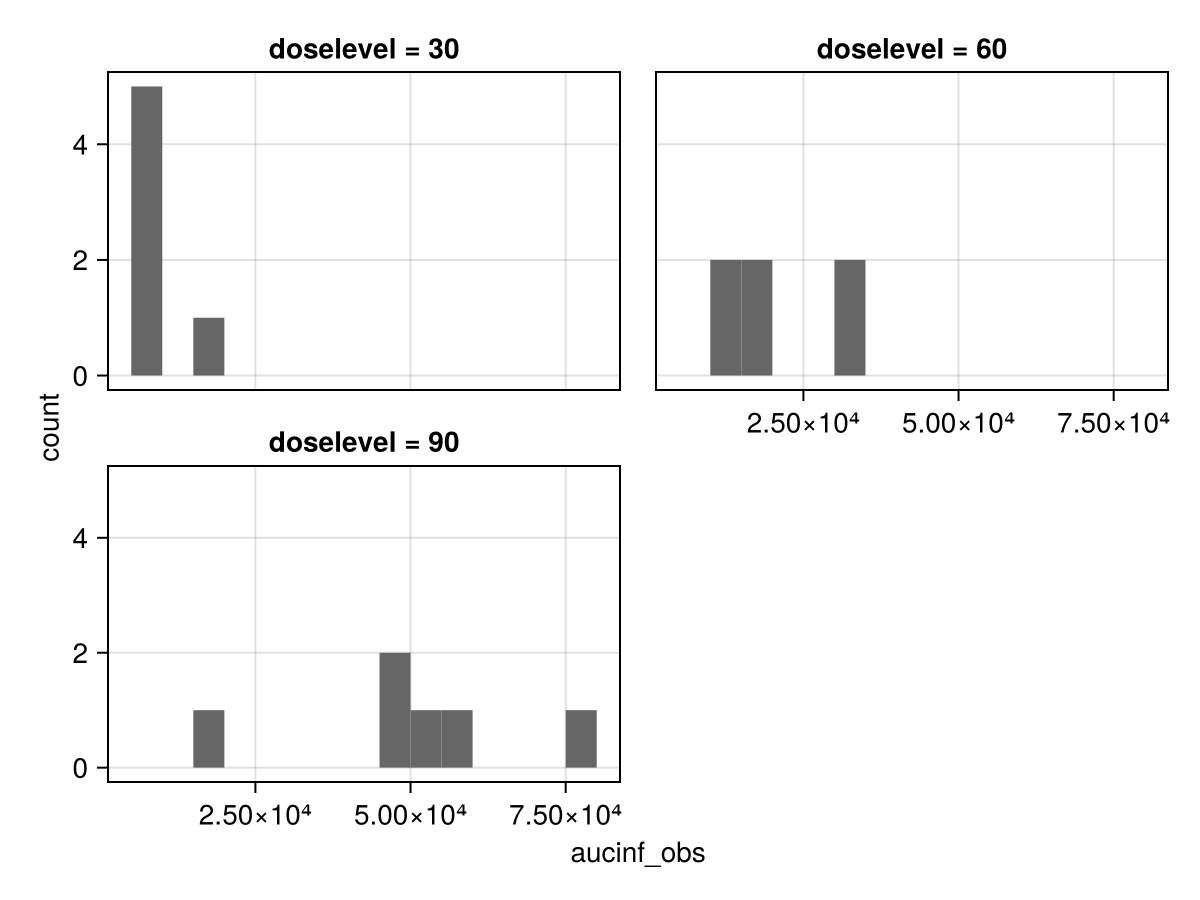
Parameter distribution by groups (ID or covariates)
parameters_vs_group(pk_nca; parameter = :aucinf_obs)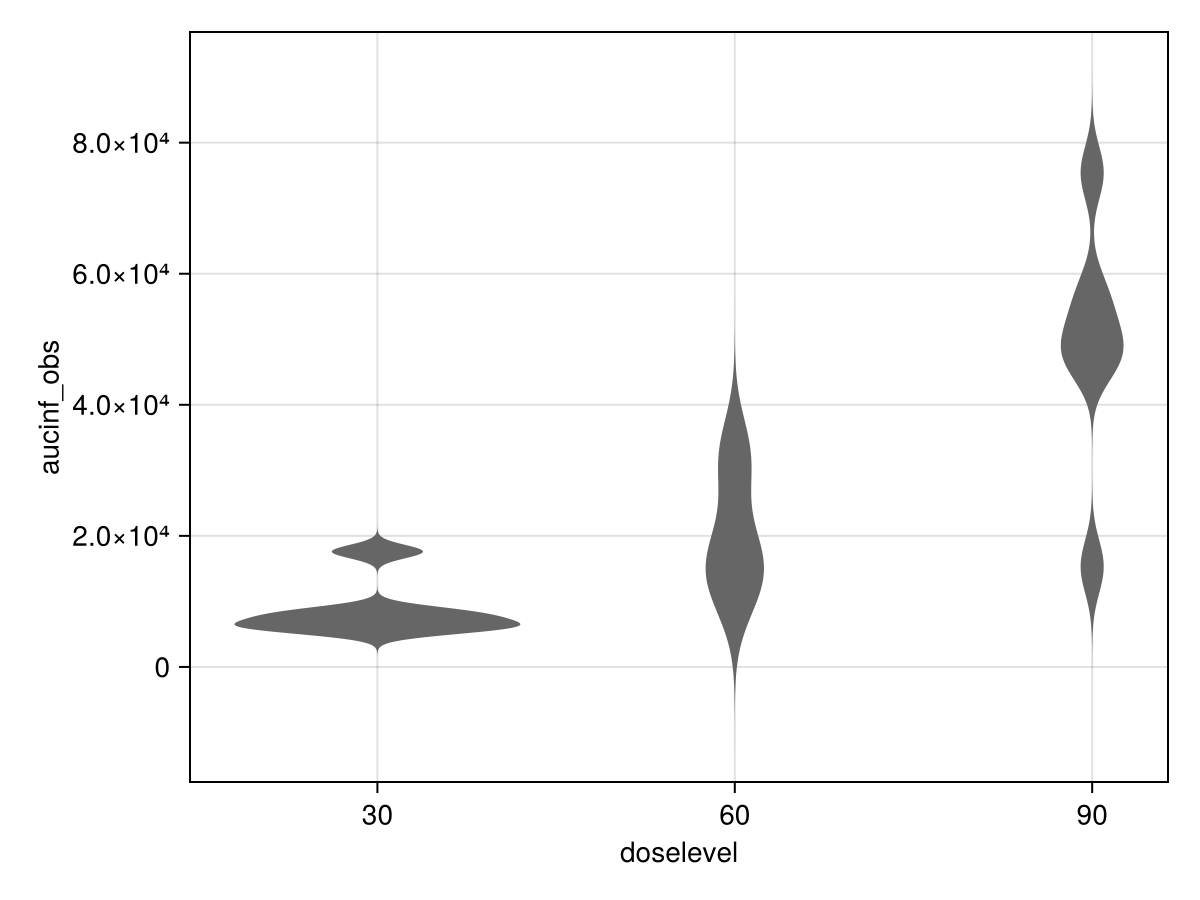
Assess dose linearity
The results of run_nca might differ significantly at different dose levels. To assess this, dose_lin fits a power regression model between dose and one of the functions described in the NCA Function List.
dose_lin = DoseLinearityPowerModel(pk_nca, :aucinf_obs; level = 0.95)Dose Linearity Power Model
Variable: aucinf_obs
Model: log(aucinf_obs) ~ log(α) + β × log(dose)
────────────────────────────────────
Estimate low CI 95% high CI 95%
────────────────────────────────────
β 1.5204 1.00121 2.03959
────────────────────────────────────The power_model function visualizes this regression model.
power_model(dose_lin)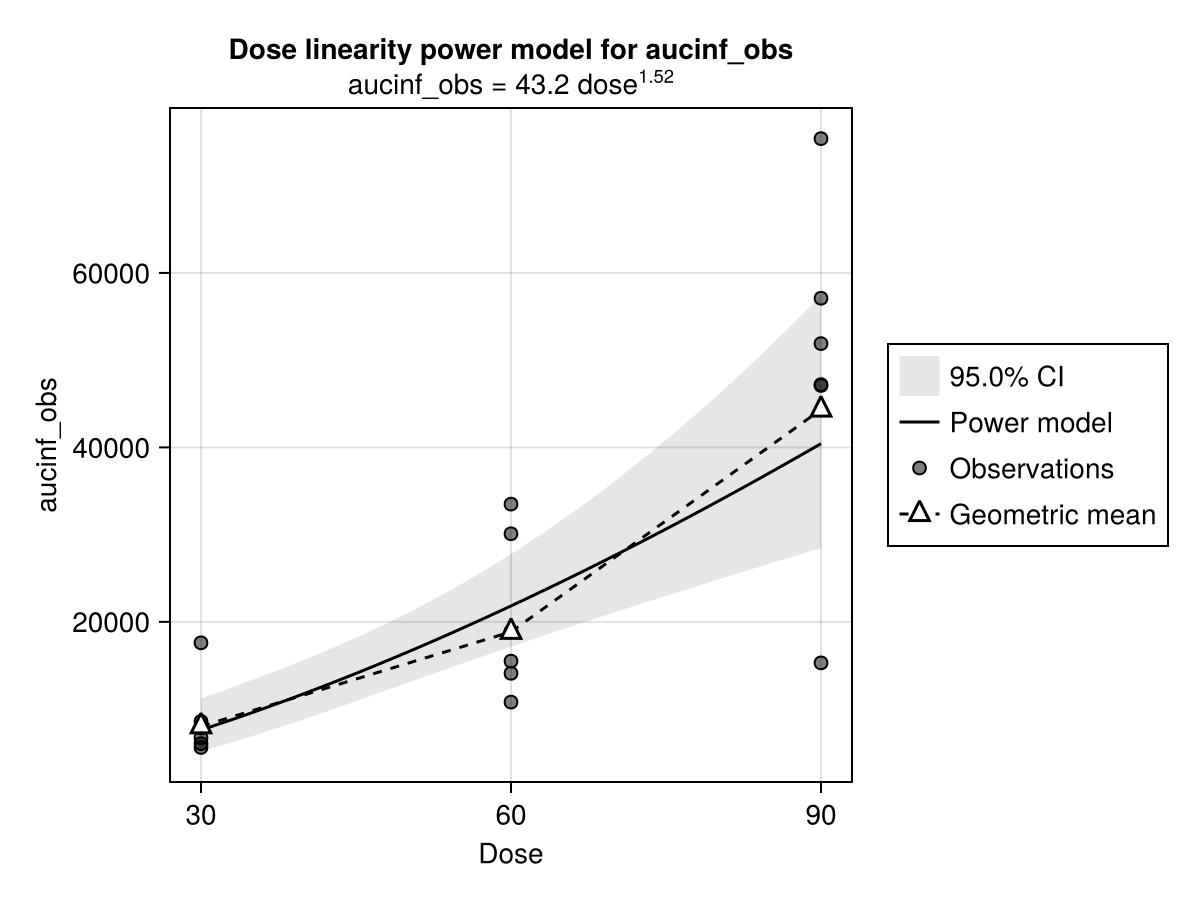
The DoseLinearityRatioTest function tests for significant differences of NCA functions between the lowest dose and the other doses.
DoseLinearityRatioTest(pk_nca, :aucinf_obs; level = 0.95)Dose linearity pairwise ratio test
Variable: aucinf_obs
───────────────────────────────────────────────────────
Ratio Estimate low CI 95% high CI 95%
───────────────────────────────────────────────────────
60.0 vs. 30.0 2.0 2.34826 1.35091 4.08194
90.0 vs. 30.0 3.0 5.52133 2.94964 10.3352
───────────────────────────────────────────────────────Generate Report
The report function takes in either a NCAPopulation or a NCAReport object to generate a comprehensive, currently only PDF, report. This generated PDF file has all the necessary information including tables, listings and figures that are required for NCA analysis. Example syntax:
report(pk_nca; output = "my_nca_report", title = "report")[ Info: Generating Tables
[ Info: Generating Plots
┌ Warning: Could not create plot.
│ error =
│ MethodError: typemax(::Type{Union{}}) is ambiguous.
│
│ Candidates:
│ typemax(::Type{T}) where T<:BitIntegers.AbstractBitSigned
│ @ BitIntegers ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/BitIntegers/xoLLM/src/BitIntegers.jl:171
│ typemax(::Type{<:Unitful.AbstractQuantity{T, D, U}}) where {T, D, U}
│ @ Unitful ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/Unitful/sxiHA/src/quantities.jl:489
│ typemax(::Type{T}) where T<:FixedPointNumbers.FixedPoint
│ @ FixedPointNumbers ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/FixedPointNumbers/Dn4hv/src/FixedPointNumbers.jl:116
│ typemax(::Type{Q}) where {T, D, Q<:DynamicQuantities.UnionAbstractQuantity{T, D}}
│ @ DynamicQuantities ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/DynamicQuantities/pNDR2/src/utils.jl:301
│ typemax(::Type{T}) where T<:BitIntegers.AbstractBitUnsigned
│ @ BitIntegers ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/BitIntegers/xoLLM/src/BitIntegers.jl:168
│ typemax(::Type{Q}) where {T, Q<:(DynamicQuantities.UnionAbstractQuantity{T})}
│ @ DynamicQuantities ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/DynamicQuantities/pNDR2/src/utils.jl:302
│ typemax(::Type{T}) where T<:Irrational
│ @ Base irrationals.jl:32
│ typemax(::Type{Q}) where Q<:(DynamicQuantities.UnionAbstractQuantity)
│ @ DynamicQuantities ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/DynamicQuantities/pNDR2/src/utils.jl:303
│ typemax(::Type{P}) where P<:Period
│ @ Dates ~/run/_work/_tool/julia/1.11.8/x64/share/julia/stdlib/v1.11/Dates/src/periods.jl:54
│ typemax(::Type{T}) where T<:IrrationalConstants.IrrationalConstant
│ @ IrrationalConstants ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/IrrationalConstants/RokwY/src/macro.jl:25
│ typemax(::Type{C}) where C<:ColorTypes.AbstractGray
│ @ ColorVectorSpace ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/ColorVectorSpace/JXxVe/src/ColorVectorSpace.jl:353
│
│ Possible fix, define
│ typemax(::Type{Union{}})
│
│ func = parameters_dist (generic function with 2 methods)
│ backtrace =
│
│ Stacktrace:
│ [1] extrema_finite(v::SubArray{Missing, 1, Vector{Missing}, Tuple{Vector{Int64}}, false})
│ @ AlgebraOfGraphics ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/AlgebraOfGraphics/RXEkR/src/scales.jl:742
│ [2] _mapreduce(f::typeof(AlgebraOfGraphics.extrema_finite), op::typeof(AlgebraOfGraphics.extend_extrema), ::IndexLinear, A::Vector{SubArray{Missing, 1, Vector{Missing}, Tuple{Vector{Int64}}, false}})
│ @ Base ./reduce.jl:437
│ [3] _mapreduce_dim(f::Function, op::Function, ::Base._InitialValue, A::Vector{SubArray{Missing, 1, Vector{Missing}, Tuple{Vector{Int64}}, false}}, ::Colon)
│ @ Base ./reducedim.jl:337
│ [4] mapreduce(f::Function, op::Function, A::Vector{SubArray{Missing, 1, Vector{Missing}, Tuple{Vector{Int64}}, false}})
│ @ Base ./reducedim.jl:329
│ [5] nested_extrema_finite(iter::Vector{SubArray{Missing, 1, Vector{Missing}, Tuple{Vector{Int64}}, false}})
│ @ AlgebraOfGraphics ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/AlgebraOfGraphics/RXEkR/src/scales.jl:746
│ [6] map(f::typeof(AlgebraOfGraphics.nested_extrema_finite), t::Tuple{Vector{SubArray{Missing, 1, Vector{Missing}, Tuple{Vector{Int64}}, false}}})
│ @ Base ./tuple.jl:355
│ [7] defaultdatalimits(positional::Vector{Any})
│ @ AlgebraOfGraphics ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/AlgebraOfGraphics/RXEkR/src/transformations/density.jl:13
│ [8] (::AlgebraOfGraphics.HistogramAnalysis{Makie.Plot{Makie.plot}, Makie.Automatic, Int64})(input::AlgebraOfGraphics.ProcessedLayer)
│ @ AlgebraOfGraphics ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/AlgebraOfGraphics/RXEkR/src/transformations/histogram.jl:71
│ [9] call_composed
│ @ ./operators.jl:1054 [inlined]
│ [10] call_composed
│ @ ./operators.jl:1053 [inlined]
│ [11] (::ComposedFunction{AlgebraOfGraphics.Visual, AlgebraOfGraphics.HistogramAnalysis{Makie.Plot{Makie.plot}, Makie.Automatic, Int64}})(x::AlgebraOfGraphics.ProcessedLayer)
│ @ Base ./operators.jl:1050
│ [12] process(layer::AlgebraOfGraphics.Layer)
│ @ AlgebraOfGraphics ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/AlgebraOfGraphics/RXEkR/src/algebra/processing.jl:184
│ [13] iterate
│ @ ./generator.jl:48 [inlined]
│ [14] collect(itr::Base.Generator{AlgebraOfGraphics.Layers, typeof(AlgebraOfGraphics.process)})
│ @ Base ./array.jl:791
│ [15] map
│ @ ./abstractarray.jl:3399 [inlined]
│ [16] AlgebraOfGraphics.ProcessedLayers(a::AlgebraOfGraphics.Layers)
│ @ AlgebraOfGraphics ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/AlgebraOfGraphics/RXEkR/src/algebra/layers.jl:54
│ [17] compute_axes_grid(d::AlgebraOfGraphics.Layers, scales::AlgebraOfGraphics.Scales; axis::@NamedTuple{})
│ @ AlgebraOfGraphics ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/AlgebraOfGraphics/RXEkR/src/algebra/layers.jl:316
│ [18] compute_axes_grid
│ @ ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/AlgebraOfGraphics/RXEkR/src/algebra/layers.jl:314 [inlined]
│ [19] #paginate#513
│ @ ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/AlgebraOfGraphics/RXEkR/src/paginate.jl:112 [inlined]
│ [20] paginate(spec::PlottingUtilities.AoGSpec; kwargs::@Kwargs{})
│ @ PlottingUtilities ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/PlottingUtilities/lT9yT/src/plots/algebraofgraphics.jl:261
│ [21] paginate
│ @ ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/PlottingUtilities/lT9yT/src/plots/algebraofgraphics.jl:257 [inlined]
│ [22] _draw_aog_spec(F::Function, args::NCAReport; paginate::Bool, kwargs::@Kwargs{figure::@NamedTuple{fontsize::Int64, size::Tuple{Int64, Int64}}, parameter::Symbol})
│ @ PlottingUtilities ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/PlottingUtilities/lT9yT/src/plots/algebraofgraphics.jl:271
│ [23] _draw_aog_spec
│ @ ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/PlottingUtilities/lT9yT/src/plots/algebraofgraphics.jl:267 [inlined]
│ [24] parameters_dist
│ @ ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/PlottingUtilities/lT9yT/src/plots/algebraofgraphics.jl:318 [inlined]
│ [25] (::NCAUtilities.var"#135#142"{NCAUtilities.var"#135#136#143"{typeof(parameters_dist), Int64, Tuple{Int64, Int64}}})(args::NCAReport; kws::@Kwargs{parameter::Symbol})
│ @ NCAUtilities ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/NCAUtilities/pf8Mo/src/reports/reports.jl:155
│ [26] report(r::NCAReport, summary::DataFrame; output::String, force::Bool, clean::Bool, header::String, footer::String, plot_fontsize::Int64, plot_resolution::Tuple{Int64, Int64}, section_order::Vector{String}, plot_styles::NCAUtilities.PlotStyles, kws::@Kwargs{title::String})
│ @ NCAUtilities ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/NCAUtilities/pf8Mo/src/reports/reports.jl:185
│ [27] top-level scope
│ @ introduction.md:229
│ [28] eval
│ @ ./boot.jl:430 [inlined]
│ [29] #59
│ @ ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/Documenter/AXNMp/src/expander_pipeline.jl:879 [inlined]
│ [30] cd(f::Documenter.var"#59#61"{Module, Expr}, dir::String)
│ @ Base.Filesystem ./file.jl:112
│ [31] (::Documenter.var"#58#60"{Documenter.Page, Module, Expr})()
│ @ Documenter ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/Documenter/AXNMp/src/expander_pipeline.jl:878
│ [32] (::IOCapture.var"#5#9"{DataType, Documenter.var"#58#60"{Documenter.Page, Module, Expr}, IOContext{Base.PipeEndpoint}, IOContext{Base.PipeEndpoint}, IOContext{Base.PipeEndpoint}, IOContext{Base.PipeEndpoint}})()
│ @ IOCapture ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/IOCapture/MR051/src/IOCapture.jl:170
│ [33] with_logstate(f::IOCapture.var"#5#9"{DataType, Documenter.var"#58#60"{Documenter.Page, Module, Expr}, IOContext{Base.PipeEndpoint}, IOContext{Base.PipeEndpoint}, IOContext{Base.PipeEndpoint}, IOContext{Base.PipeEndpoint}}, logstate::Base.CoreLogging.LogState)
│ @ Base.CoreLogging ./logging/logging.jl:524
│ [34] with_logger(f::Function, logger::Base.CoreLogging.ConsoleLogger)
│ @ Base.CoreLogging ./logging/logging.jl:635
│ [35] capture(f::Documenter.var"#58#60"{Documenter.Page, Module, Expr}; rethrow::Type, color::Bool, passthrough::Bool, capture_buffer::IOBuffer, io_context::Vector{Any})
│ @ IOCapture ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/IOCapture/MR051/src/IOCapture.jl:167
│ [36] runner(::Type{Documenter.Expanders.ExampleBlocks}, node::MarkdownAST.Node{Nothing}, page::Documenter.Page, doc::Documenter.Document)
│ @ Documenter ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/Documenter/AXNMp/src/expander_pipeline.jl:877
│ [37] dispatch(::Type{Documenter.Expanders.ExpanderPipeline}, ::MarkdownAST.Node{Nothing}, ::Vararg{Any})
│ @ Documenter.Selectors ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/Documenter/AXNMp/src/utilities/Selectors.jl:170
│ [38] expand(doc::Documenter.Document)
│ @ Documenter ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/Documenter/AXNMp/src/expander_pipeline.jl:60
│ [39] runner(::Type{Documenter.Builder.ExpandTemplates}, doc::Documenter.Document)
│ @ Documenter ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/Documenter/AXNMp/src/builder_pipeline.jl:224
│ [40] dispatch(::Type{Documenter.Builder.DocumentPipeline}, x::Documenter.Document)
│ @ Documenter.Selectors ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/Documenter/AXNMp/src/utilities/Selectors.jl:170
│ [41] #90
│ @ ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/Documenter/AXNMp/src/makedocs.jl:283 [inlined]
│ [42] withenv(::Documenter.var"#90#92"{Documenter.Document}, ::Pair{String, Nothing}, ::Vararg{Pair{String, Nothing}})
│ @ Base ./env.jl:265
│ [43] #89
│ @ ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/Documenter/AXNMp/src/makedocs.jl:282 [inlined]
│ [44] cd(f::Documenter.var"#89#91"{Documenter.Document}, dir::String)
│ @ Base.Filesystem ./file.jl:112
│ [45] makedocs(; debug::Bool, format::Documenter.HTMLWriter.HTML, kwargs::@Kwargs{plugins::Vector{CitationBibliography}, modules::Vector{Module}, doctest::Bool, warnonly::Vector{Symbol}, clean::Bool, draft::Bool, linkcheck::Bool, linkcheck_timeout::Int64, linkcheck_ignore::Vector{Regex}, sitename::String, authors::String, remotes::Nothing, pages::Vector{Any}})
│ @ Documenter ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/Documenter/AXNMp/src/makedocs.jl:281
│ [46] top-level scope
│ @ ~/run/_work/PumasDocs.jl/PumasDocs.jl/docs/make.jl:72
│ [47] include(mod::Module, _path::String)
│ @ Base ./Base.jl:562
│ [48] exec_options(opts::Base.JLOptions)
│ @ Base ./client.jl:316
│ [49] _start()
│ @ Base ./client.jl:524
└ @ NCAUtilities ~/run/_work/PumasDocs.jl/PumasDocs.jl/custom_julia_depot/packages/NCAUtilities/pf8Mo/src/reports/reports.jl:162
[ Info: Gathering System Information
[ Info: Ordering Report SectionsYou can view the report generated here.